UMCD
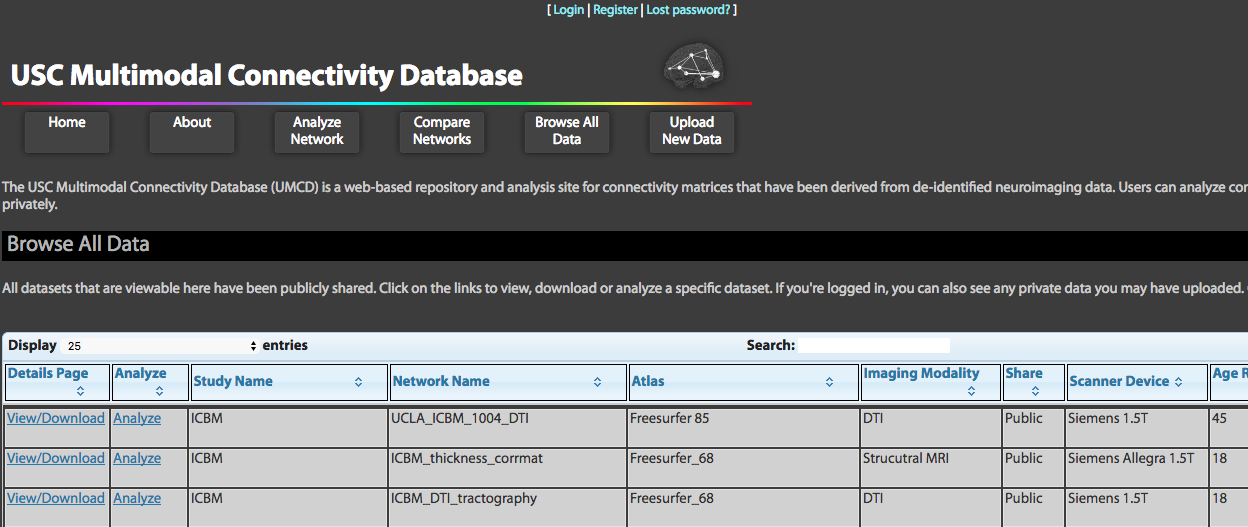
The USC Multimodal Connectivity Database (UMCD) is a web-based repository and analysis site for connectivity matrices that have been derived from de-identified neuroimaging data. Users can analyze connectivity matrices that have been shared publicly and upload their own matrices to share or analyze privately.
Modulzyer
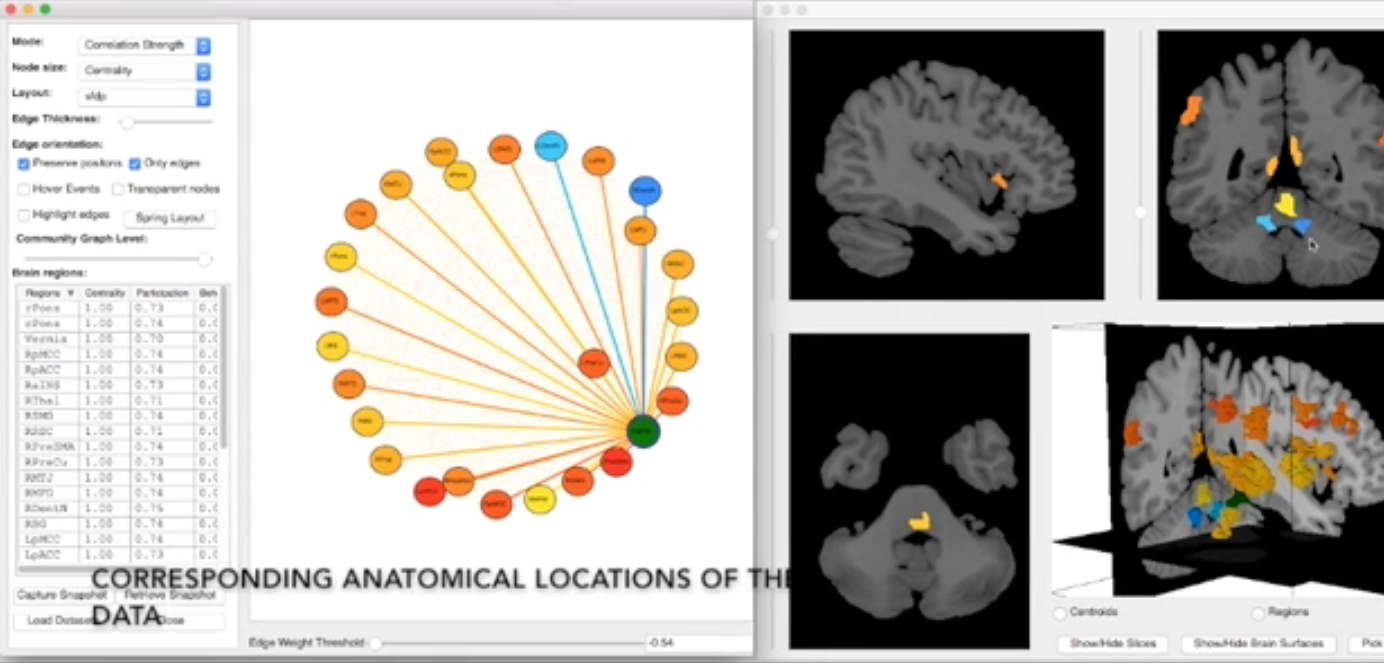
Brain Modulyzer is an interactive visual exploration tool for functional magnetic resonance imaging (fMRI) brain scans, primarily developed by Sugeerth Murugesan. It is aimed at analyzing the correlation between different brain regions when resting or when performing mental tasks. Brain Modulyzer combines multiple coordinated views—such as heat maps, node link diagrams and anatomical views—using brushing and linking to provide an anatomical context for brain connectivity data. Integrating methods from graph theory and analysis, e.g., community detection and derived graph measures, makes it possible to explore the modular and hierarchical organization of functional brain networks. Providing immediate feedback by displaying analysis results instantaneously while changing parameters gives neuroscientists a powerful means to comprehend complex brain structure more effectively and efficiently and supports forming hypotheses that can then be validated via statistical analysis.
UMCP
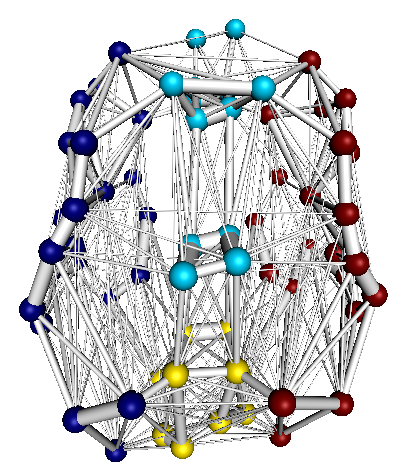
UMCP is a set of Python programs used to calculate connectivity matrices and metrics from a variety of neuroimaging modalities including diffusion weighted MRI (DTI/DSI; tracks.py), fMRI (timeseries.py), and structural MRI. It will also generate 2D and 3D network visualizations (plot_network.py).